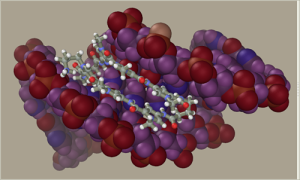
Improved interface with NWChem.
Increased speed of molecular dynamics.
Fixed bugs.
Ascalaph v. 1.8.54
Posted in Uncategorized
Ascalaph v. 1.7.11
Improving the stability of long simulations.
Posted in Uncategorized
Ascalaph v. 1.7.10
Localization.
Speed improvement.
Posted in Uncategorized
Ascalaph v. 1.7.8
General сleaning version
Posted in Uncategorized
Ascalaph
I would like to introduce a program of molecular modeling Ascalaph
at http://www.biomolecular-modeling.com/Ascalaph/index.html
General Features:
- Molecular graphics
- Molecular model building
- Geometry optimization
- Molecular dynamics
- Quantum modelling using NWChem, CP2K and PC GAMESS/Firefly
- Parallel molecular dynamics on Linux clusters with MDynaMix molecular dynamics package
- Hardware-accelerated simulations using NVIDIA CUDA technology
Much attention is paid to design the initial models. Molecules can be drawn by hand, then put in the minimal reasonable conformation with aid of rough optimization, and then optimized by quantum-chemical methods. There is a constructor of chain molecules, which allows to create a model with pre-installed molecular-mechanical parameters form the residues. Users can create their own residues and add them to the database. There is a procedure for adding the box with a solvent, such as a water. This box should be made separately and then added to the model. The overlapping solvent molecules can be automatically removed.
Interface with quantum chemical programs primarily designed to develop the parameters of molecular mechanical force fields. Currently supported NWChem, CP2K and PC GAMESS/Firefly.
I will be very grateful for comments and suggestions.
Posted in Molecular modeling | Tags: Biology, Chemistry, Force Field, molecular, molecular modeling, Monte Carlo, Physics, Quantum, Research, simulation, Software